Assembling the Tree of Life (ATOL) program funded by the US NSF fund aims at reconstruction the evolutionary history of several millions of species. This goal cannot be achieved by contemporary ordinary phylogenetic reconstruction methods. This calls for a supertree approach – an approach for combining small trees over partial, overlapping, sets of species, into a big tree over the complete species set.
Quartet trees – trees over four species, are the most basic informative phylogenetic unit as it can define uniquely any tree. Therefore, quartet amalgamation – joining quartets into a single tree lies at the heart of almost any phylogenetic task. Quartet supertree arises in many instances, even when the history is not tree-like, with event like horizontal gene transfer or incomplete lineage sorting. Nevertheless, it is among the hardest computationally and basic questions are open for decades.
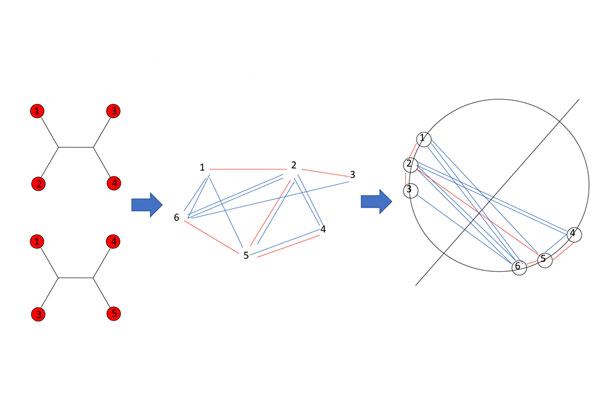
During the years, we have developed a novel graph-based, divide and conquer technique relaying on semi-definite programming (SDP) approach. The approach, denoted Quartet MaxCut, constructs the Quartet Graph based on the input quartets, and then applies a MaxCut to this graph. We have devised a super fast implementation to this approach that is the fastest and most accurate for this task. Our software was used by many users in many applications ranging from cancer detection and even up to computer graphics and music classification.
In parallel, in a series of theoretical works, we could show that the MaxCut approach yields the first theoretical approximation guarantee, of 0.425, improving over the naïve 1/3 obtained trivially by a random tree.
- Snir and S. Rao. Quartet MaxCut: A Fast Algorithm for Amalgamating Quartet Trees. Molecular Phylogenetics and Evolution, (MPE), 2012.
- Snir and S. Rao. Quartets MaxCut: A Divide and Conquer Quartets Algorithm. IEEE/ACM Transactions on Computational Biology and Bioinformatics (TCBB), Volume 7(4): 704-718 (2010).
- Avni*, Z. Yona*, R. Cohan, S. Snir. The performance of Two Supertree Schemes Compared Using Synthetic and Real Data Quartet Input. Journal of Molecular Evolution. *Authors contributed equally.
- Avni*, R. Cohen*, and S. Snir. Weighted Quartets Phylogenetics. Systematic Biology. 64 (2): 233-242, 2015. *Authors contributed equally.
- Snir and R. Yuster. Reconstructing approximate phylogenetic trees from quartet samples. SIAM Journal on Computing (SICOMP). 41(6): 1466-1480 (2012) . Extended version of ACM-SIAM Symposium on Discrete Algorithms (SODA) 2010.
- Alon, S. Snir and R. Yuster. On the compatibility of quartet trees, SIAM J. Discrete Math (SIDMA). 28(3), 2014. Extended version of ACM-SIAM Symposium on Discrete Algorithms (SODA) 2014.
- Snir and R. Yuster. A Linear Time Approximation Scheme for Maximum Quartet Consistency on Sparse Sampled Inputs. SIAM J. Discrete Math. 25(4): 1722-1736, 2011. Extended version of The International Workshop on Approximation Algorithms for Combinatorial Optimization (APPROX) 2011.